The long and ...long of it:
|
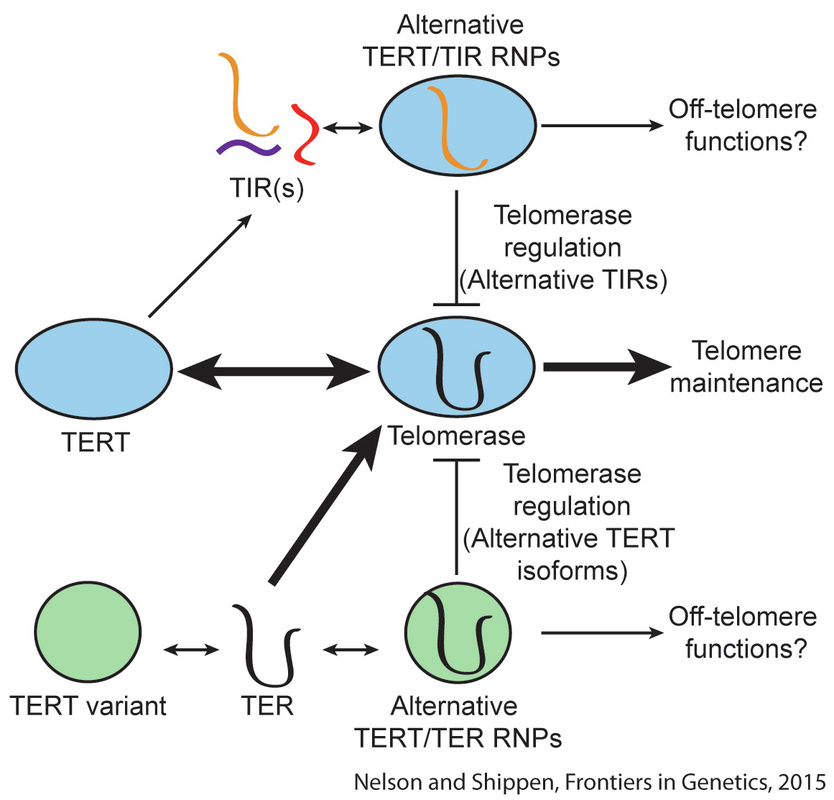
Long non-coding RNAs (lncRNAs) comprise a large, enigmatic portion of eukaryotic transcriptomes. The functions of most lncRNAs are still poorly understood, but the lncRNAs that have been described perform diverse roles covering the breadth of cell development, metabolism, and maintenance. I first became aware of the importance of lncRNAs during my graduate research on telomerase regulation in Dorothy Shippen’s lab at Texas A&M. Telomerase is a ribonucleoprotein complex found in most eukaryotes that maintains the ends of chromosomes, thus ensuring the complete transfer of genetic material from parent to offspring. One of the core telomerase constituents is a lncRNA known as TER (see right). While I was in Dorothy’s lab, two TERs were identified in Arabidopsis thaliana. I determined, both in vitro and in vivo, that the second TER, TER2, inhibits telomerase activity during periods of intense genotoxic stress (1). TER2’s telomerase inhibition is facilitated by the incorporation of a transposable element into the TER2 locus (2). Thus, TER2 is actually a TERT-interacting RNA (TIR) that acts as both an environmental sensor (of DNA damage) and a molecular sponge, preventing TERT from illegitimately adding telomeric DNA at double strand breaks (see right). From both a functional and an evolutionary perspective, TER2 represents but one example of how lncRNAs can rapidly evolve to take on complex regulatory roles in fundamental biological pathways.
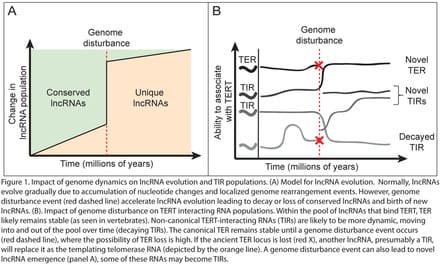
TERs are just one example of the different lncRNA functions described to date. In a broad sense, lncRNAs can be divided into four functional categories: a) chromatin-dependent scaffolds, such as HOT-AIR and XIST; b) RNAs involved in transcriptional regulation and mRNA splicing, such as ASCO-RNAs in Arabidopsis and MALAT1 in mammals; c) molecular decoys, such as the alternative telomerase RNA (TER2) in A. thaliana or the regulation of apoptosis by lncRNA Gas5; and d) chromatin-independent scaffolds, such as TER in eukaryotes. In summary, lncRNAs regulate a huge range of processes, including cell and organismal development, genomic stability, epigenetic regulation, and response to environmental cues (3). For an excellent set of reviews, see Ulitsky and Bartel (2013); Ponting et al, (2009); Ariel et al, (2015); and Wang and Chang (2011). While many of the currently identified lncRNAs may be transcriptional “noise,” there are still likely thousands of lncRNAs that remain uncharacterized. One of my goals in Mark Beilstein’s lab at the University of Arizona has been to develop an evolutionary transcriptomics framework by which I can identify conserved lncRNAs and then predict function based on conserved motifs. An important finding from my work is that plant lncRNAs emerge and decay more rapidly than in mammals, and much of this variation can be attributed to genomic perturbation events that occur regularly in plants. Acquiring the computational skills necessary to perform my comparative analyses led me to collaborate with Eric Lyons and the iPlant consortium at the University of Arizona. Through transcriptomic and genomic analyses in the plant family Brassicaceae, I identified ~1,200 A. thaliana lncRNAs that are conserved and expressed across >43 million years of evolution (4). Overlaying genome and lncRNA annotation datasets has allowed me to infer potential regulatory functions for many of these RNAs. A large proportion of these conserved lncRNAs are either stress-responsive (drought, cold, or heat), or contain micro RNA binding motifs. Thus, these lincRNAs represent a very interesting group of molecules that need further functional studies. |