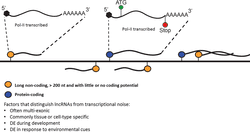
The first way that we distinguish any transcript is by trying to fit it into a category. So how do we define lncRNAs? Well, lncRNAs look a lot like mRNAs in that many are Pol-II transcribed, capped and tailed, and can be multi-exonic. Now, in an attempt to filter lncRNAs from transcriptional noise, we can also limit the distinguisher “lncRNA” to transcripts that show tissue or cell-type specific expression, or to transcripts that are differentially expressed during development or in response to environmental cues. These specific patterns in expression are usually our only hints that these transcripts are functional lncRNAs.
What do they do?
Classification of lncRNAs have coalesced around four main functional archetypes: molecular signals, decoys, guides, and scaffolds (K., C., Wang and Chang, 2011; Ulitsky and Bartel, 2013). As a molecular signal, lncRNAs serve as rapid responders to diverse cellular stimuli. Molecular signals can perform their function through the mere act of transcription or through interactions with protein-binding partners (Pontier and Gribnau, 2011; Hawkes et al., 2016; Vance and Ponting, 2014). Molecular decoy lncRNAs act as sinks for a broad range of RNA-binding proteins, likely through RNA-structure dependent interactions (Hansen et al., 2013; Redon et al., 2010). Guide lncRNAs recruit proteins to specific locations along a chromosome, thus requiring both lncRNA-protein and lncRNA-DNA interactions (Lee, 2009). Lastly, scaffold lncRNAs serve to recruit multiple binding partners into a single functional ribonucleoprotein complex, irrespective of chromosomal context (Zhang et al., 2011).
Specific examples: Mammalian MALAT1
MALAT1 is a 8708 nt long highly expressed lncRNA that serves as a prognostic marker for multiple cancers and enhances tumor growth by promoting proliferation, invasion, and metastasis. It is believed to perform at least part of these processes by altering gene expression and mRNA splicing. MALAT1 is also necessary for normal cell growth, making it difficult discern “normal” from “abnormal” MALAT1 functions. While MALAT1 levels are upregulated in cancer cells, it is already a highly abundant nuclear lncRNA, suggesting that the regulatory pathway(s) where MALAT1 functions is quite complex.
Eukaryotic Telomerase RNA component (TERC)
For eukaryotes, passing on a complete set of genetic material to their offspring is made difficult by the biochemistry underlying DNA replication. Normal DNA replication is unable to extend the ends of chromosomes, which we call telomeres. This causes telomeres to shorten each round of cell division until at some point, in multi-cellular organisms, the cell stops dividing, going into a senescent stage of their life cycle where they drop out of the pool of dividing cells. In order to pass on the entire genome to offspring, eukaryotes needed a way to extend and maintain telomeres. Most eukaryotic cells manage chromosome end maintenance by a complex called telomerase, which is comprised of a reverse trancriptase TERT, a lncRNA subunit TERC, and a variety of subunits that assist in biogenesis of the complex and activity. TERC is important here not only as a scaffold to bring these proteins together, but also because it contains a small template domain with sequence similarity to the telomere. This template domain allows TERT to add telomeric sequence to the end of the chromosome.